File Sink
Used to write a stream to a binary file.
This file can be read into any programming environment that can read binary files (MATLAB, C, Python, ...). It can also be played back in GRC using a File Source. For example, if complex type is chosen, then the binary file will be full of float32s in IQIQIQ order. There is no meta data or anything else included with the binary data. For more information on handling this data, see the #Handling_File_Sink_data section below.
Parameters
(R): Run-time adjustable
- File (R)
- Path of the file to open and write output to. If the specified file name does not exist at that location, it creates a file of that name over there. Otherwise, if the file already exists, it may overwrite or append the file based on the append option.
- Unbuffered
- Specifies whether the output is buffered in memory. If the output is unbuffered, the data will be flushed to the file each time the work function is called. This can cause the flowgraph to run slow due to the time required to access the disk each time.
- Append File
- Gives an option to either append to the file or to overwrite the file.
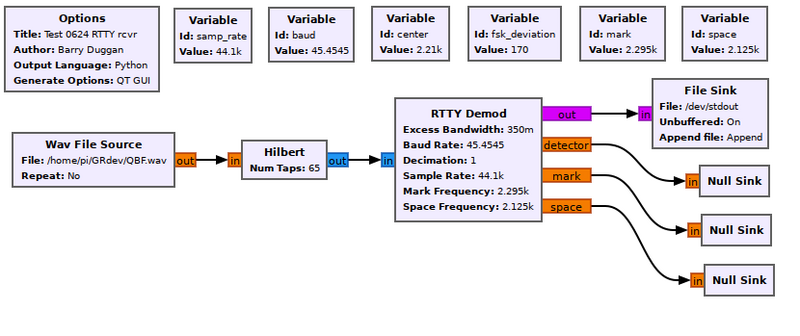
Example Flowgraph
This flowgraph shows a File Sink which outputs text to the terminal (/dev/stdout).
Handling File Sink data
What is the file format of a file_sink? How can I read files produced by a file sink?
All files written by File Sink are in pure binary format. Just bits. That's it. A floating point data stream is saved as 32 bits in the file, one after the other. A complex signal has 32 bits for the real part and 32 bits for the imaginary part. Reading back a complex number means reading in 32 bits, saving that to the real part of a complex data structure, and then reading in the next 32 bits as the imaginary part of the data structure. And just keep reading the data.
Take a look at the Octave and Python files in gr-utils for reading data using Octave and Python's Scipy module.
The exception to the format is when using the metadata file format. These files are produced by the File Meta Sink block and read by the File Meta Source block. See the manual page on the Metadata Information for more information about how to deal with these files.
Reading from Python
A one-line Python command to read the entire file into a numpy array is:
import numpy
f = numpy.fromfile(open("filename"), dtype=numpy.uint8)
Replace the dtype with numpy.int16, numpy.int32, numpy.float32, numpy.complex64 or whatever type you were using.
Reading from Octave / Matlab
If you're using octave (or are stuck with using Matlab, our condolences), the following is a method to read all floats written with a File Sink to a variable:
f = fopen('filename', 'rb');
values = fread(f, Inf, 'float');
Replace 'float' with 'short','int' or 'char' as appropriate.
Use complex_v = values(1,:) + values(2,:)*i; to convert interleaved real, imaginary values to an array of complex values.
Reading from plain C
This is the easiest, since we assume you know how `fread` works if you're writing C, and how to cast a pointer. So, here's a way of simply mapping your file to memory and use that to access the values, directly. This is sensible if you do not intend to load the full file into RAM, but want to jump around in it.
// this is C99.
#include <sys/mman.h>
#include <sys/stat.h>
#include <stdio.h>
#include <complex.h>
…
int fd = open("filename", "rb"); // please check fd != -1
struct stat stat_struct;
int retval = fstat(fd, &stat_struct); // please check for ==0
size_t filesize = stat_struct.st_size;
char* address = mmap(NULL, filesize, PROT_READ, MAP_PRIVATE, fd, 0); // please check for address != MAP_FAILEP
// interpret as pointer to a complex float value (could alternatively use float*, int*, ...
complex float* complex_values = (complex float*) address;
// do stuff on complex_values[i]...
…
// finally, clean up:
munmap(address, filesize);
close(fd);
Source Files
- C++ files
- TODO
- Header files
- TODO
- Public header files
- TODO
- Block definition
- TODO